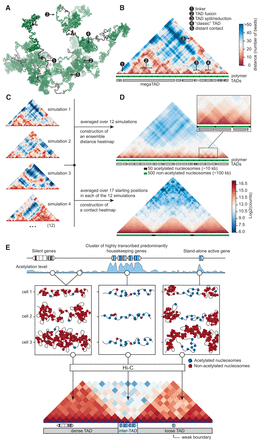
Computer simulation of a linear polymer that folds into a set of TADs supports a key role of hyperacetylated chromatin in separation of TADs. (A) One of the predicted spatial configurations of a polymer composed of 19 blocks of inactive (interacting) nucleosomes (500 nucleosomes each, green) interspaced by shorter blocks of active noninteracting nucleosomes (50 nucleosomes each, black). (B) Spatial proximity map (distance heat map) of the polymer configuration presented in A. Distances are measured in numbers of nucleosomes. A scheme of the model polymer and positions of TADs predicted by the Armatus algorithm are shown below the map. (C) Distance heat maps of the four individual configurations of the model polymer. Configuration 1 is used in A and B. (D) Distance heat map (upper) and contact heat map (simulated Hi-C map constructed at resolution of 4 kb [20 beads] for three consecutive TADs, lower) of the model polymer obtained by the averaging of the heat maps of 12 individual configurations. Notation as in B. (E) A schematic illustrating the proposed model of chromatin folding into TADs/inter-TADs, as directed by the self-association of nucleosomes. A high acetylation level of the chromatin within the genomic regions harboring actively transcribed genes interferes with chromatin packaging into TADs due to decreased inter-nucleosomal interactions.











