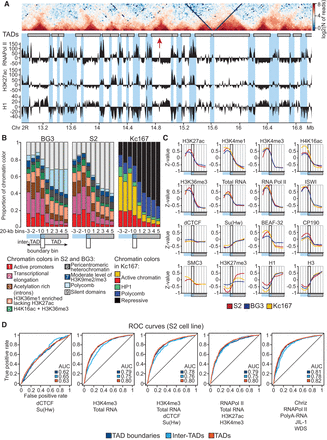
Partitioning of chromosomes into TADs and inter-TADs reflects distributions of active and repressed chromatin regions. (A) Heat map of a 4-Mb region of Chromosome 2R aligned with modENCODE tracks of RNA polymerase II, H3K27ac, and histone H1 for BG3 cells. TAD boundaries and inter-TAD regions are shaded in blue. Red arrow indicates an unidentified weak TAD boundary. (B) Distribution of chromatin types (colors) near TAD boundaries. Box plots show proportions of chromatin colors in bins located at the same position relative to a TAD boundary, averaged over all TADs. TAD boundaries were aligned so that inter-TAD bins were placed to the left of the boundary bin (bin 0) and TAD bins were placed to the right of the boundary bin. The region from the third bin in the inter-TAD (left) to the fifth bin in the TAD (right) is shown. Plots with longer distances from the boundary are shown in Supplemental Figure S5A. Chromatin colors are taken from Filion et al. (2010) for the Kc167 cells and from Kharchenko et al. (2011) for the S2 and BG3 cells. The OSC cells were not analyzed, as epigenetic data were not available for this cell line. The P-values are presented in Supplemental Table S6. (C) Distribution of the individual chromatin marks and proteins near TAD boundaries. Curves smoothed with LOESS show the median Z-transformed values in the groups of bins described above. Thick rectangles show TAD boundary bins. Box plots are shown in Supplemental Figure S6. P-values are presented in Supplemental Table S6. (D) Prediction of TADs, inter-TADs, and TAD boundaries in the S2 cells using logistic regression models based on the genome distribution of active chromatin marks or architectural proteins dCTCF and Su(Hw). Receiver operating characteristics (ROC curves) and AUC (area under the curve) values are shown.











