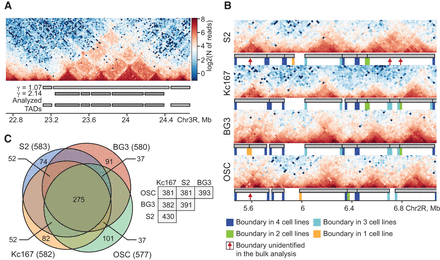
Genomic positions of topologically associating domains (TADs) are largely conserved among Drosophila cells of different origins. (A) A fragment of the Hi-C interaction map (heat map) for the BG3 cell line. Gray rectangles below the heat map represent TADs predicted with two different values of the scaling parameter γ. This illustrates a two-step TAD prediction method utilizing a higher γ value for large TADs containing internal self-interacting domains. (B) Heat maps of a 1.6-Mb region of Chromosome 2R with annotated TADs. Positions of boundary bins are indicated by colored rectangles under the TADs map. The color code for the boundary bins is shown at the bottom. Red arrows indicate unidentified weak boundaries. The resolution of all heat maps is 20 kb. (C) Venn diagram showing the numbers of TADs shared between all studied cell lines (with both boundaries located at the same genomic bin or at adjacent bins). Total numbers of TADs in each cell line are shown in parentheses. Numbers of TADs shared between different pairs of the cell lines are shown to the right of the diagram.











