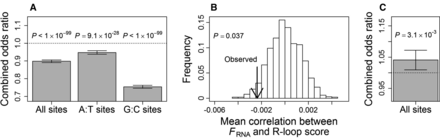
Human population genomic data of introns reveal the negative impact of nascent RNA folding strength (FRNA) on R-loop formation and mutation rate. SNPs are considered to have higher mutation rates than non-SNP control sites. (A) Evidence for a negative impact of FRNA on mutation rates at all sites, A/T sites, and G/C sites. Odds ratio <1 indicates that SNP sites have lower FRNA than non-SNP control sites within a gene. Shown are the results combined from all genes by the MH procedure. Error bars represent 95% confidence intervals estimated by bootstrapping the genes 1000 times. The dotted line indicates an odds ratio of 1. (B) The among-gene average of the within-gene Pearson's correlation between FRNA and R-loop score is significantly more negative than the random expectation. The arrow indicates the actual observation, whereas the bars show the frequency distribution of the corresponding value derived from 1000 sets of genes with FRNA values randomly shuffled within genes. (C) Evidence for a positive impact of R-loop on mutation rate. Odds ratio >1 indicates that SNP sites have higher R-loop scores than non-SNP control sites within a gene. Shown are the results combined from all genes by the MH procedure. Error bars represent 95% confidence intervals, estimated by bootstrapping the genes 1000 times. The dotted line indicates an odds ratio of 1.











