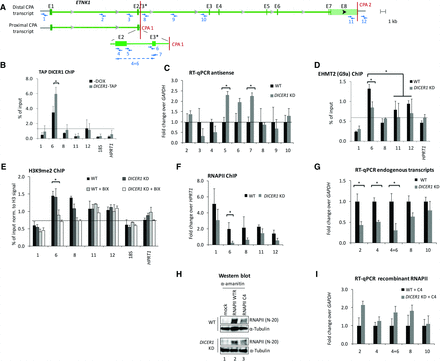
DICER1 regulates proximal CPA site usage at the ETNK1 locus. (A) Schematic showing positions of ChIP amplicons and RT-qPCR probes (1–12) on the human ethanolamine kinase 1 (ETNK1) locus around proximal and distal CPA sites 1 and 2 (CPA 1 and 2): (E3*) alternative exon 3. (B) ChIP analysis of TAP-tagged DICER1 occupancy over ETNK1. Primer pairs analyze the region surrounding ETNK1 CPA sites: (6, 8) intronic CPA 1; (11, 12) canonical CPA 2. Values from noninduced and DICER1 KD HEK293 cells are shown as a percentage of input. Mean values and standard deviations were calculated from at least three biological replicates: (*) P < 0.05 by two-tailed Student's t-test. (C) Antisense RNA levels around ETNK1 at CPA 1. Primer pairs specific to intronic and exonic antisense transcripts were used to determine relative changes in antisense RNA levels upon DICER1 KD by RT-qPCR. (D,E) ChIP analysis as in B of EHMT2 histone methyltransferase (D) and histone 3 lysine 9 dimethylation (H3K9me2) (E): (18S) 18S ribosomal RNA; (BIX) BIX-01294 trihydrochloride hydrate. (F) RNA polymerase II (RNAPII) occupancy at human ethanolamine kinase 1 (ETNK1) CPA sites. The percentage of input values from noninduced and DICER1 KD HEK293 cells are shown as fold change over HPRT1 signals. (G,I) Levels of intronic and exonic sense transcripts around ETNK1 CPA 1 from endogenous RNAPII transcription (G) and upon reconstitution of transcription with recombinant RNAPII construct C4 (I). Signals from noninduced and DICER1 KD HEK293 cells are shown as fold over wild type and normalized to GAPDH sense transcript levels. Mean values and standard deviations were calculated from at least three biological replicates: (*) P < 0.05 by two-tailed Student's t-test. (H) Western blot showing expression levels of recombinant, α-amanitin resistant RNAPII mutant WTR (recombinant wild type) and C4 (slow elongating mutant) in wild-type and DICER1 KD HEK293 cells: (mock) empty vector.











