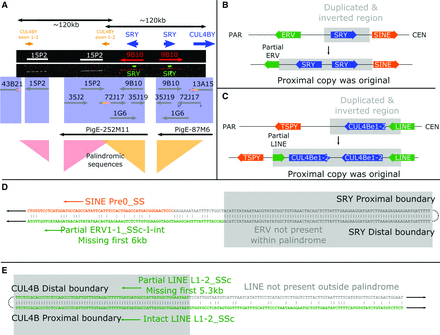
The pig SRY region. The Yp proximal block of genes contains two overlapping palindromes of ∼120 kb each. These surround the duplicated sequences CUL4BY exons 1–2 and SRY. (A) FISH results from Y fosmid clones and probes for the SRY gene are shown with the BAC and fosmid clone sequences found mapping to the region. The inversion boundaries are both identifiable; the CUL4BY inversion runs from the last 3 kb of 43B21 to within 72J17; the SRY inversion begins also within 72J17 and runs to 13A15. A schematic view is also shown of the regions surrounding the SRY (B) and the CUL4B duplications (C). The SRY duplication disrupts an ERV element, revealing the proximal copy to be ancestral. The CUL4B duplication copies part of a LINE element, again revealing the proximal copy to be ancestral. The sequence alignments across the inversion breakpoints are shown in more detail for SRY (D) and CUL4B (E). The order of events was therefore a duplication of SRY, including the first two exons of CUL4B, followed by duplication of the region around the CUL4B copy.











