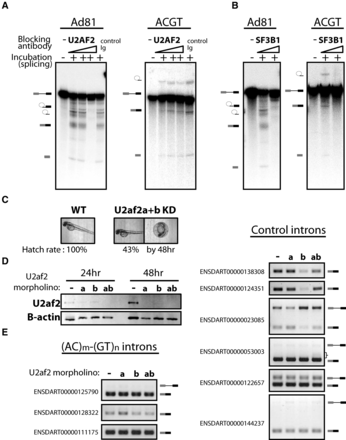
The splicing of (AC)m-(GT)n introns requires components of U2 snRNP but not U2AF2. In vitro splicing substrates were prepared from the constructs used in Figure 4 and the model in vitro splicing construct Ad81. The in vitro–transcribed RNA was incubated in HeLa nuclear extract for the splicing assay. (A) The splicing of the Ad81 control was compared to the (AC)m-(GT)n intron with pre-incubation with either: no antibody, a control antibody, or increasing amounts of anti-U2AF2 antibody. The input pre-mRNA and the splicing products, including the lariat intermediate and free first exon from the first step of splicing, the free lariat, and ligated exon from the second step of splicing, were resolved on an 8M urea gel and visualized by autoradiography with a phosphoimager. (B) The comparison described above was repeated with antibody targeting SF3B1, a component of the U2 snRNP. (C) u2af2 knockdown (KD) in zebrafish embryos. The KD embryos developed without gross phenotypic defect by 48 h but with lower hatch rate. (D) Western blotting of U2af2 24 or 48 h after injection. Beta actin served as a loading control. (E) RT-PCR of single intron transcripts containing (AC)m-(GT)n repeats (left) or without the repeats (right). Pre-mRNA and/or spliced mRNA is depicted on the right of the gel images. A bracket indicates the smear of the PCR product, possibly due to the loss of splicing accuracy.











