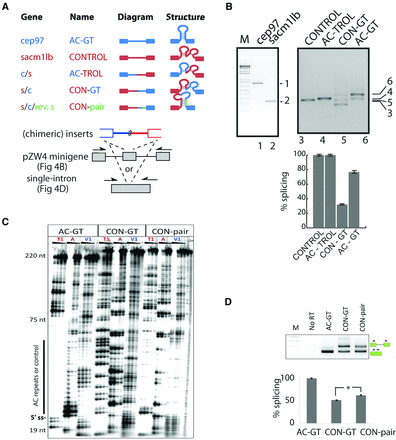
The predicted hairpin in (AC)m-(GT)n introns is required for accurate splice site recognition. (A) The exemplar (AC)m-(GT)n intron from zebrafish cep97 intron 5 (blue) and a control from sacm1lb intron 16 (red) were recombined at their midpoints and cloned into splicing reporters to make chimeric intronic constructs. Construct nomenclature derives from suffix/prefix combinations listed on the left. Green color indicates regions of complementarity used to make compensatory mutations in CON-pair constructs. (B) RT-PCR from total RNA extracted from whole-body juvenile zebrafish. Primer amplicons span four exons centered on the AC-GT or CONTROL intron. RT-PCR from total RNA extracted from transiently transfected tissue culture cells and quantified (histogram). The predicted sizes of the constitutively spliced products are illustrated with arrows marked by lane numbers. (C) RNase mapping of AC-GT, CON-GT, and CON-pair introns. Structured regions inferred from protection from single-strand nucleases (see RNases T1 and A). (D) Intron substrates used in mapping were tested in a single intron splicing reporter construct, transfected into cells, and assayed and quantified as described above. Asterisk indicates statistical significance by paired t-test (P = 9 × 10−4) of three biological replicates.











