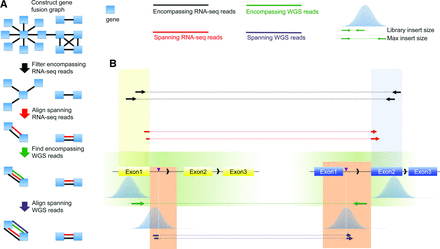
Overview of INTEGRATE. (A) INTEGRATE establishes a gene fusion graph using encompassing RNA-seq reads (black lines) to connect nodes or genes (blue rectangles). Edges of the fusion graph are removed following various filtering steps before undergoing a targeted split-read alignment involving the remaining edges. Encompassing and spanning split-read realignment and mapping are performed on BWTs of gene nodes (Supplemental Fig. 1). Encompassing WGS reads are retrieved from regions determined by spanning RNA-seq reads. Spanning WGS reads are aligned to the regions indicated by encompassing WGS reads (steps indicated by green and purple arrows; also see B). (B) When encompassing (black) and spanning (red) RNA-seq reads have been mapped to the genes involved in a gene fusion, the encompassing WGS reads (green) are expected from focal encompassing WGS regions (green area) bounded by maximum insert size upstream of or downstream from the fusion junctions of the transcripts. The spanning WGS reads (purple) are expected to align within focal WGS regions (orange area) bounded by fusion junction and maximum insert size downstream from the encompassing WGS reads.











