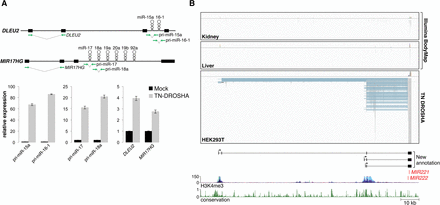
DROSHA inhibition facilitates pri-miRNA assembly. (A) qPCR analysis of pri-miRNA abundance in HEK293T cells with or without expression of TN-DROSHA. The assayed transcripts DLEU2 and MIR17HG are depicted in the top panel with green arrows indicating the location of primers. qPCR results are shown in the bottom panel with error bars representing standard deviations derived from three independent measurements. (B) Visualization of RNA-seq data from Illumina Human BodyMap 2.0 (kidney and liver) and TN-DROSHA-transfected HEK293T cells. The Integrative Genomics Viewer (IGV) was used to visualize mapped read alignments. Segments of reads that are aligned to the genome are shown in gray, while blue lines represent spliced sequences. StringTie assembled transcripts produced from this locus are shown at the bottom of the panel. Plots representing H3K4me3 histone marks and evolutionary conservation were generated using the UCSC Genome Browser (human genome GRCh37/hg19 assembly). The y-axes for UCSC Genome Browser tracks shown in this and all other figures represent the default vertical viewing range settings.











