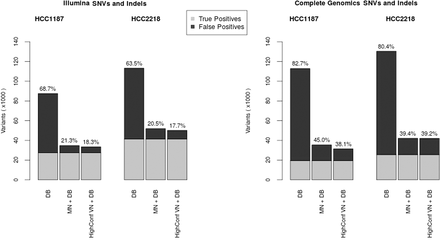
Number of false-positive SNVs and indels identified per filtering method for Illumina (left) and Complete Genomics (right). True positives (light gray) are those variants remaining after application of all filters (for VN, we did not use the high-confidence criterion to determine the set of true positives). DB denotes an aggressive database filter. HighConf VN+DB denotes the list of high-confidence somatic variants as determined by the VN and database filters. MN + DB denotes the list of high-confidence somatic variants after correction with a MN combined with the database filter.











