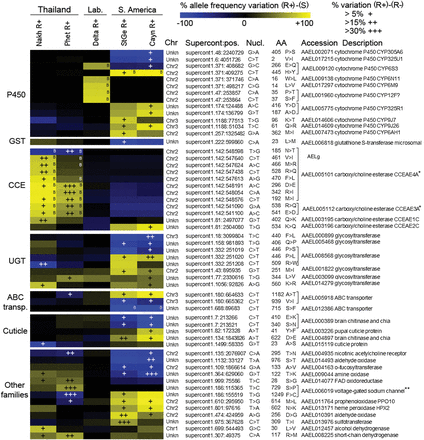
Best nonsynonymous polymorphisms associated with deltamethrin resistance. Only the 55 best differential nonsynonymous variants identified from frequency-based filtering and the Bayesian approach are shown (see Methods). For each region, allele frequency variation between each resistant population (R+ phenotypes) and their susceptible counterpart (S) are shown as a blue-yellow color scale. Blue indicates an enrichment in the reference allele, whereas yellow indicates an enrichment in the variant allele. Variants identified by the Bayesian approach are indicated by “B” marks. For variants passing frequency-based filtering or BayeScan3 filtering, allele frequency variation between R+ and R− phenotypes are shown as overimposed “+” marks. Variants are grouped by gene families and are described by the following annotations: chromosomal location (according to Juneja et al. 2014; Timoshevskiy et al. 2014), supercontig position, nucleotide change, amino acid position, amino acid change, gene accession number, and gene description. (*) Genes also found affected by CNVs linked to deltamethrin resistance. (**) The sodium channel S729P and F1249C variants correspond to the S989P and F1534C kdr mutations described in the literature due to changes in AAEL006019-RD transcript annotation.











