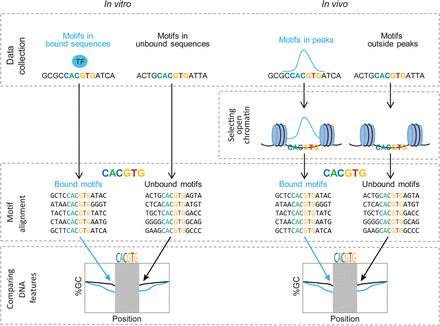
A flowchart describing the approach used for finding preferences in regions surrounding TF binding motifs. (Left) For each TF, a pool of bound and unbound sequences was collected from HT-SELEX data of human and mouse TFs (Jolma et al. 2013). Both sequence pools were filtered, keeping only sequences possessing the published TF binding motif. The sequences were further aligned relative to the TF binding motif. Nucleotide content of the sequences flanking the binding motif was compared between the bound and unbound groups. (Right) For each TF, a pool of bound and unbound sequences was collected from ChIP-seq data (Yan et al. 2013). Both pools were filtered, keeping only sequences possessing the TF binding motif in open chromatin. The sequences were further aligned relative to the motif. Finally, the nucleotide content of the sequences surrounding the binding motif in the bound and unbound groups was compared between the two groups.









