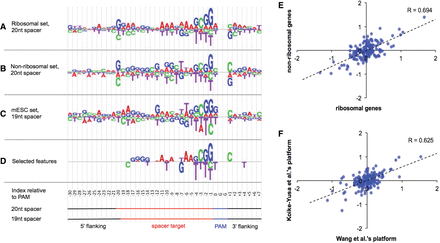
Preference of nucleotide sequences that impact sgRNA efficiency. (A–C) Logos showing the sequence preference of the three sgRNA sets defined in Figure 1. The height of the nucleotides represents the log odds ratio of nucleotide frequency between efficient and inefficient sgRNAs. (D) A logo showing the selected features that reproducibly impact sgRNA efficiency in the three sgRNA sets. The height of the nucleotides represents the coefficients computed from the Elastic-Net. (E,F) Scatter plots showing the correlation of sequence preference for sgRNAs targeting ribosomal versus nonribosomal genes in Wang data (E) and sgRNAs in Wang data versus Koike-Yusa data (F). Each dot represents a nucleotide in a 40-bp region centered by the spacer. The sequence preference is measured as the log2 odds ratio of nucleotide frequency between efficient and inefficient sgRNAs.











