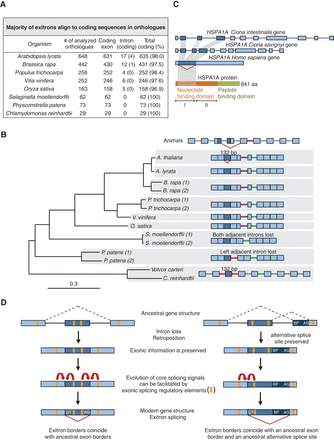
Origin and evolution of EIs. (A) Statistics of EI alignments to plant orthologous sequences. (B) EI evolution in the gene encoding glycine cleavage system T-protein. Phylogenetic reconstruction of intron loss events in different plant species (1 and 2 indicate paralogs). Gene structures are not to scale. Introns at the conserved positions are colored. EI and the homologous sequences are in dark blue. Red carets, EIS and ES events. (C) EI evolution in the intronless human HSPA1A gene. The human EI corresponds to three coding exons in Ciona spp. (D) Evolution of EIS, the “splicing memory” hypothesis. Evolution of a subset of EIs involved loss of introns and retroposition. Upon intron loss, exonic information was preserved. If ancestral regions were subjected to alternative splicing (dashed lines), vestigial exonic splicing regulatory elements present close to the former exon borders could facilitate evolution of core splicing signals and the reestablishment of an AS event in the modern gene by EIS.











