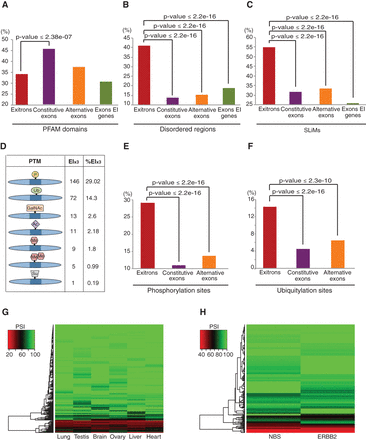
Functional implications of EIS in human. (A–C) Enrichment of PFAM domains (A), disordered regions (B), and SLiMs (C) in protein sequences encoded by EIs and other types of exons. (D) Statistics of various post-translational modification (PTM) sites encoded by EIs. (E,F) Enrichment of phosphorylation (E) and ubiquitylation (F) sites in the sequences encoded by EIs and other types of exons. (A–F) Analyses performed for EIx3 subset only. (G,H) Heatmap of EIS (measured by PSI) in different human tissues (G) and in ERBB2-positive breast cancer and normal breast tissue (NBS) samples (H).











