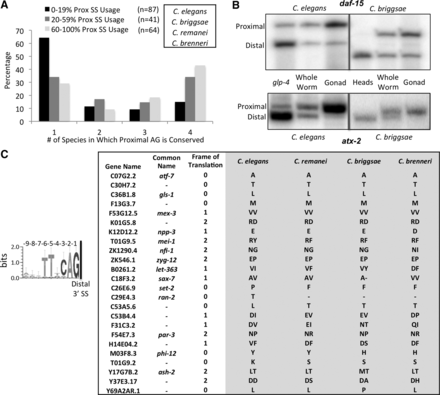
Adjacent alternative 3′ splice sites are conserved in related Caenorhabditis species. (A) Bar graph depicting conservation of the proximal AG dinucleotide in tissue-specific alternative 3′ splice sites. Introns were separated according to percent of proximal 3′ splice site usage in the wild-type germline (0%–19% in black, 20%–59% in dark gray, and 60%–100% in light gray). The comparison is made with three species related to C. elegans: C. remanei, C. briggsae, and C. brenneri. Displayed is a bar graph depicting the percentage of events in each category in which the upstream AG dinucleotide is conserved in only C. elegans (1) or in C. elegans plus one, two, or three other related species (2, 3, and 4). (B) 32P-labeled RT-PCR products made from primers that flank introns in daf-15 and atx-2 with tissue-specific alternative adjacent 3′ splice sites. Enrichment in C. elegans or C. briggsae for somatic cells (glp-4 or dissected heads, respectively) and germline cells (dissected gonads) shows a shift in abundance from the distal site isoform to the proximal site isoform in the germline of both species. (C) WebLogo consensus sequence motif of the 9 nt preceding the distal 3′ splice site of the 53 introns with tissue-specific 3′ splice sites separated by 9 nt (from Supplemental Table S2). Note the lack of conserved nucleotides at positions −7 to −9 from the distal 3′ splice site. On the right, a table depicting the 24 introns with the AG dinucleotide at the proximal site 9 nt upstream of the distal splice site conserved in all four nematode species analyzed. Shown is the gene name, common name, the frame of translation, and the identity of the amino acid(s) derived from the −7 to −9 nt.











