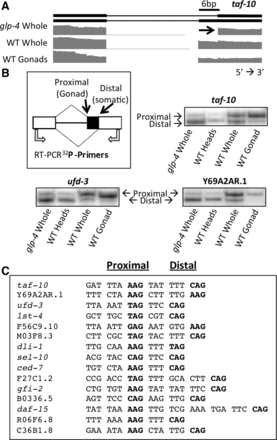
The major class of splicing changes between germline and somatic cells consists of adjacent alternative 3′ splice sites. (A) taf-10 alternative isoforms and RNA sequencing coverage tracks showing a tissue-specific alternative 3′ splicing event. Germline reads primarily cross the 3′ splice site closer to the 5′ splice site (proximal) while reads in somatic cells primarily use the 3′ splice site further from the 5′ splice site (distal). (B) Cartoon depicting the proximal and distal 3′ splice sites, the tissue-type in which each 3′ splice site is primarily used (in parentheses), and the location of the 32P-labeled oligos used to validate the tissue-specific enrichment of each isoform by RT-PCR. 32P RT-PCR products from three sample genes were separated on a 6% polyacrylamide denaturing gel and visualized with a PhosphorImager. (C) Nucleotides preceding and within tissue-specific 3′ splice sites in a representative set of genes. The proximal and distal 3′ splice sites are in bold for each sequence (left and right, respectively). Nucleotides are spaced to show the 3-nt periodicity and maintenance of frame between the 3′ splice sites.











