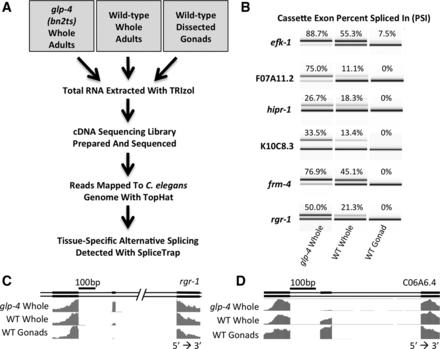
Cassette exon alternative splicing changes between C. elegans germline and somatic tissues. (A) Flowchart depicting the process of RNA isolation, high-throughput sequencing, and computational analysis. (B) RT-PCR products from primers that flank alternative cassette exons highlighting tissue-specific inclusion/skipping. Products were separated on an Agilent 2100 Bioanalyzer. PSI was calculated by dividing the molarity of the inclusion product by the sum of the molarities of the inclusion and skipping products. (C) Representation of the WormBase gene annotation for rgr-1 alternative isoforms and normalized RNA sequencing coverage tracks for the indicated libraries. This demonstrates skipping of the alternative cassette exon in gonad. (D) Representation of the WormBase gene annotation for C06A6.4 showing alternative isoforms and RNA sequencing coverage. This demonstrates an example of a cassette exon that is specifically included in the germline.











