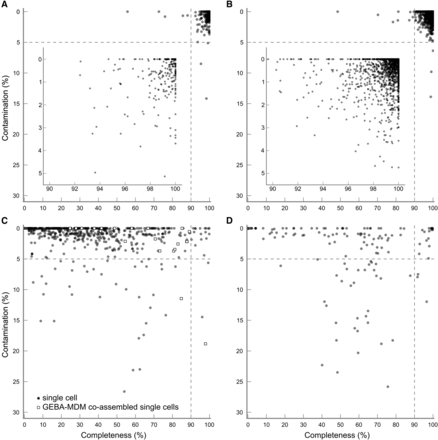
Lineage-specific completeness and contamination estimates for 262 isolates annotated as finished in IMG (A), 2019 isolates annotated as draft in IMG (B), 632 genomes recovered using single-cell genomics (C), and 146 population genomes recovered from metagenomic data (D). Dashed lines indicate the criteria required for a genome to be considered a near-complete genome with low contamination. Insets give a more detailed view of the quality of the isolate genomes. The 2281 isolate genomes were obtained from IMG and sequenced as part of the GEBA, GEBA-KMG, GEBA-PCC, GEBA-RNB, or HMP initiatives.











