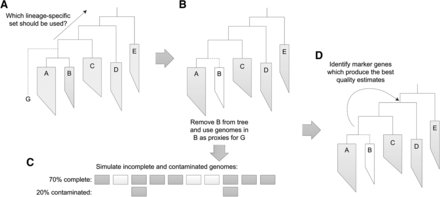
Overview of simulation framework for selecting lineage-specific marker genes. (A) To evaluate a genome, G, it is placed into a reference genome tree. Each parental node from the point of insertion to the root of the tree defines a lineage-specific marker set which can be used to estimate the completeness and contamination of this genome. (B) To select a suitable set of lineage-specific marker genes for evaluating G, the genomes in the child lineage of G with the fewest genomes were used as proxies for G. (C) Genomes at different levels of completeness and contamination were simulated from these proxy genomes by subsampling and duplicating fixed sized genomic fragments. (D) Each parental marker set was then used to estimate the completeness and contamination of these simulated genomes, and the marker set resulting in the best average performance over all simulated genomes was identified. This marker set is used to assess the quality of any genomes subsequently inserted along this branch.











