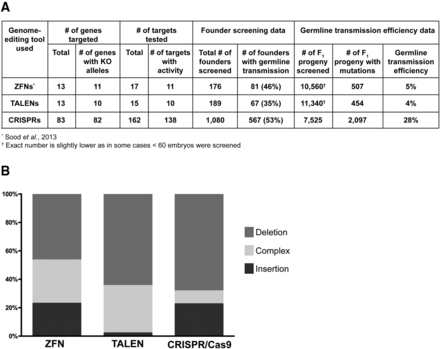
Summary and comparison of CRISPR/Cas9 to ZFN- or TALEN-induced mutations. (A) Summary of mutant identification data compiled from CRISPR/Cas9, TALENs, and ZFNs. The mutants were identified using fluorescent PCR and the aggregate data from each technique is shown. (B) Comparison of the types of mutations detected by sequencing. The mutations were classified in three different groups: deletions, insertions, and complex (at least one deletion and at least one insertion detected within ±30 bp of the 3′ end of the sgRNA or ZFN/TALEN target).











