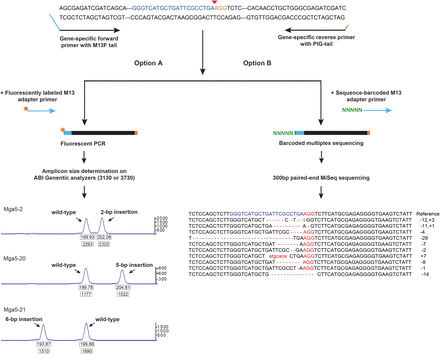
Overview of the mutant identification strategies. Mutant alleles were identified by fluorescence PCR and high-throughput sequencing. Gene-specific primers were designed to amplify the regions around the target site (200–300 bp). The forward primer contains an M13-forward sequence on the 5′ end, and the reverse primer contains a pig-tail sequence to reduce size heterogeneity PCR artifacts. Depending on the application, one of two additional primers was added. (Option A) For fluorescent PCR, an M13-forward oligo with the fluorophore FAM attached was added. The resulting primers were run on an ABI 3100 or 3730 along with a size standard to obtain the amplicon sizes accurate to base-pair resolution. The size of the peak relative to the wild-type peak control determines the nature and length of the mutant. (Option B) When amplicons were sequenced, a third primer containing M13-forward sequence with a 6-bp barcode attached to the 5′ end was used. A unique barcode was assigned to each embryo from the same target. The amplicons were sequenced on an Illumina MiSeq platform with 300-bp paired-end reads. The sequences were processed using ampliconDIVider for the identification of insertions and deletions induced by CRISPR/Cas9 (http://research.nhgri.nih.gov/CRISPRz).











