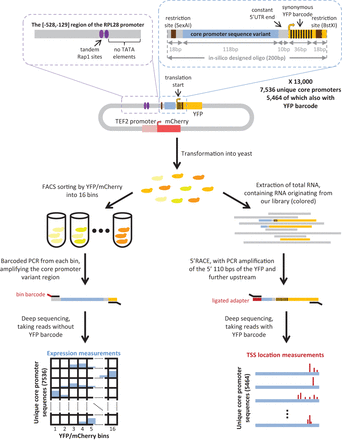
Illustration of our experimental system. Oligonucleotides from a library comprising 13,000 designed synthetic sequences (Agilent Technologies) containing a 118-bp-long variable core promoter sequence were ligated into a low copy plasmid (top). Designed sequences included 7536 unique core promoter sequences, and for 5464 of them, we designed a second instance that was additionally barcoded by synonymous mutations within the first 36 bp of the YFP. The barcoded sequences also differed from the nonbarcoded ones by four mismatches within the 10 bp upstream of the YFP. The plasmid pool was transformed into yeast to create a heterogeneous pool of yeast cells, each cell expressing YFP at a different level (middle). To measure expression, cells were sorted using fluorescence activated sorting (FACS) into 16 expression bins by their YFP/mCherry ratio, and the core promoter sequences were amplified using bin-specific barcoded primers and sent to parallel sequencing (left pipeline). Sequencing reads coming from YFP-barcoded instances were removed. Each read was then mapped to a YFP/mCherry bin and a core promoter sequence. This gave for each core promoter sequence the binned distribution of YFP/mCherry levels over the cells that had that sequence (bottom left), from which we computed the mean YFP/mCherry (see Supplemental Note). To map TSSs, we extracted total RNA from the pool of yeast cells, performed 5′ RACE using primers specific to the YFP sequence, and sequenced the products (right pipeline). Sequencing reads not mapping to YFP-barcoded instances were removed. Each read was then mapped to a core promoter sequence by its YFP barcode, enabling us to compute the transcription initiation landscape of YFP-barcoded core promoter sequences (bottom right) (see Supplemental Note).











