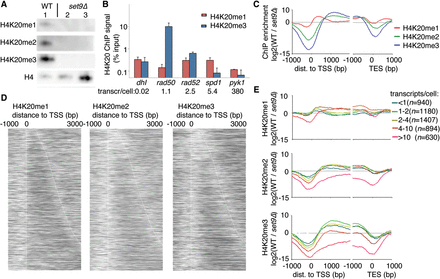
H4K20 methylation is found in protein-coding regions of the genome. (A) Western blot analysis of wild-type (WT) cells (lane 1) and set9Δ cells (lanes 2,3 differ in the amount of chromatin loaded). (B) ChIP-qPCR analysis of H4K20me1 and H4K20me3 at five loci that have different transcription levels. Bars show the average and SEM of three independent experiments. (C) The average pattern of H4K20 methylation marks aligned with the TSS (left) or TES (right). (D) Patterns of H4K20me1 (left), H4K20me2 (middle), and H4K20me3 (right) aligned at the TSS, with genes sorted according to length (top, short genes; bottom, long genes). (E) Metagene analysis of the levels of H4K20me1 (top), H4K20me2 (middle), and H4K20me3 (bottom), stratified according to gene expression level.











