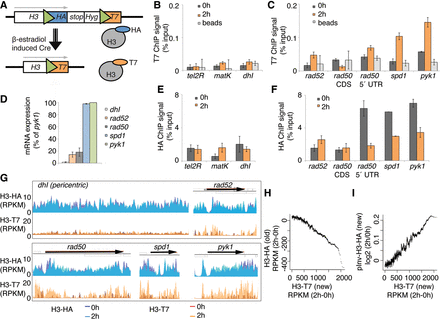
The RITE system that was used to measure genome-wide nucleosome turnover in S. pombe. (A) Model of the RITE genetic switch system. Green triangles represent LoxP sites. (B,C) T7 ChIP-qPCR analysis of heterochromatic regions (B) and genes (C). (D) RNA levels at five loci in growth-arrested (3 h at 36°C) cells with the RITE system (Hu2549 with cdc25-22 allele). (E,F) HA ChIP-qPCR of heterochromatin regions (E) and genes (F). Three independent experiments were performed, and the error bars show SEM. (G) A browser view of HA and T7 ChIP-exo results for five genomic loci before (0 h) and after (2 h) the genetic switch. Data show the average signal from two independent experiments. (H) The running average (window 500) of ChIP-exo signals of 150-bp chromatin fragments from H3-HA versus H3-T7 (RITE). (I) The running average (window 500) of ChIP-exo signals of 150-bp chromatin fragments from H3-T7 (RITE) versus pInv-H3-HA.











