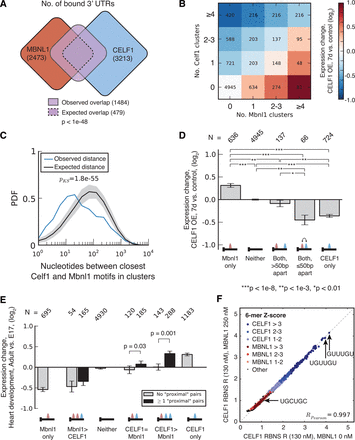
CELF1 and MBNL1 bind in close proximity to the same 3′ UTRs and exert opposing effects on mRNA stability. (A) Venn diagram showing the expected and observed overlap between CELF1 and MBNL1 3′ UTR targets (CELF1 data from muscle, MBNL1 data from myoblasts). The observed overlap is approximately three times larger than expected (analysis controlled for gene expression) and significant by Fisher's exact test. (B) Expression change following CELF1 induction in muscle (7 d versus control) for transcripts grouped by number of MBNL1 and CELF1 CLIP clusters in the 3′ UTR. CLIP clusters for MBNL1 and CELF1 were derived from myoblast and muscle, respectively. (C) The probability density function (PDF) of the distribution of distances between CELF1 CLIP clusters in muscle and MBNL1 CLIP clusters in myoblasts, in 3′ UTRs with binding for both proteins. Distances for true binding sites are shown in blue; for randomly placed binding sites, in black. Statistical significance was assessed by modified KS test, and the distribution of distances in shuffled controls is shown in gray. (D) Expression change following CELF1 induction in muscle (7 d versus control) for transcripts with exactly one MBNL1 CLIP cluster and exactly one CELF1 CLIP cluster. Genes were grouped according to whether the distance between motifs is less than or greater than 50 base pairs (bp). Significance was assessed by rank-sum test. (E) Expression change during heart development (adult versus E17) for transcripts with varying numbers of MBNL1 and CELF1 CLIP clusters. Genes were grouped according to the relative abundance of MBNL1 and CELF1 CLIP clusters and by the presence/absence of a proximal MBNL1/CELF1 binding pair. Only transcripts with gene expression MZ scores >0.5 were included in this analysis, and significance was assessed by KS test. See also Supplemental Figures S7 and S8 and Supplemental Tables S7 and S8. (F) MBNL1 presence does not affect in vitro binding of CELF1 to RNA. 130-nM tagged CELF1 and 250-nM MBNL1 were equilibrated in vitro with random RNA 40mers in an RNA Bind-n-Seq experiment. The tagged CELF1 was pulled down, bound 40mers were eluted and sequenced, and RBNS R of each 6mer was calculated as the frequency in the pulldown library divided by the frequency in the input library. 6mers were classified as weak, medium, or strong (RBNS R Z-score between 1–2, 2–3, or >3, respectively), CELF1-binding, MBNL-binding, or none of the above (“Other”) using the data from Lambert et al. (2014) and color-coded as indicated.











