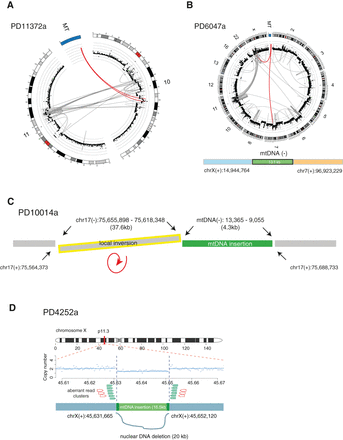
Concurrence of somatic mtDNA nuclear transfers with other structural variations. The complex web of rearrangements in the vicinity of mitochondrial-nuclear DNA fusions from four examples. (A) In PD11372a, mtDNA integration with complex rearrangements between Chr 10 and 11. (B) In PD6047a, mtDNA integration with complex rearrangements among Chr 6, 7, 11, 22, and X. (A,B) DNA copy numbers are shown by black dots with a log scale. Red lines represent translocations involving mtDNA. (C) In PD10014a, mtDNA integration combined with a local inversion (yellow). (D) In PD4252a, mtDNA integration with a local deletion. DNA copy numbers are shown with blue dots and lines. Aberrant read clusters (discordant and split reads) are shown by green and red arrows, respectively.











