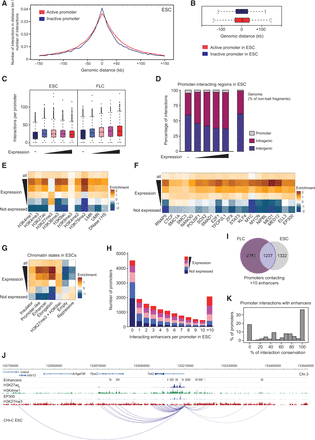
Hallmarks of promoter-interacting regions. (A) Composite profile showing the proportion of promoter–genome interactions for 5-kb distance bins upstream of and downstream from the transcription start sites for active (red) and inactive (blue) promoters in ESCs. (B) Genomic range of interactions for active (red) and inactive (blue) promoters in ESCs. (C) Number of promoter–genome interactions in ESCs and FLCs, separated by expression categories. (D) Intra- and intergenic distribution of promoter-interacting regions in ESCs, with genes driven by the promoters separated in expression categories (HindIII fragments encompassing exonic or intronic sequences are classed as “intragenic” here). The distribution of intragenic and intergenic sequences in the mouse genome is shown on the right. (E–G) Heat maps showing the enrichment/depletion for histone modifications (E), chromatin proteins (F), and chromatin states (G) in promoter-interacting regions in ESCs, for all promoters and separated by expression of the interacting promoters, compared to nonbait regions. (UMR) Unmethylated region; (LMR) low-methylated region (Stadler et al. 2011). (H) Number of promoters from each expression category interacting with between zero and more than 10 genomic elements with the hallmarks of enhancers in ESCs. (I) Unique and overlapping promoters interacting with multiple (>10) enhancer-like elements in ESCs and FLCs. (J) Example of a promoter (driving the Tet2 gene) contacting multiple enhancers in ESCs. (K) Conservation of promoter–enhancer contacts between ESCs and FLCs. Shown is the percentage of promoters that share 0%, 10%, 20%, etc., of their interactions with enhancer-like elements. Only enhancers active in both cell types (ESC and FLC) have been included in the analyses.











