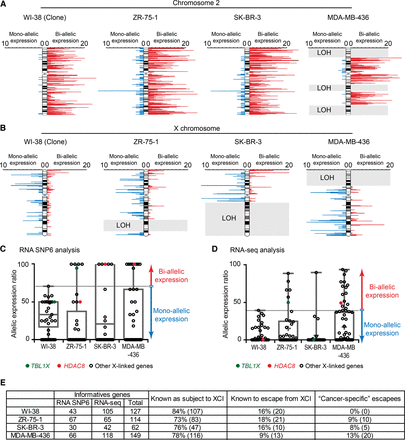
Abnormal reactivation of the inactive X chromosome in breast cancer cell lines. (A,B) RNA SNP6 analysis shows the expression status of an autosomal chromosome, as example Chromosome 2 (A), and the X chromosome (B) in normal (WI-38) and breast cancer cell lines (ZR-75-1, SK-BR-3, and MDA-MB-436). Red bars indicate biallelic expression, and blue bars indicate monoallelic expression. The bar length represents the number of expressed informative SNPs on a 50-SNP sliding window. Gray rectangles correspond to noninformative regions due to loss of heterozygosity (LOH). Two WI-38 subclones (#1 and #28), carrying an inactive X chromosome of opposite parental origin, show clear monoallelic expression from either the maternal or paternal X chromosome confirming the clonality of the subclones (see Supplemental Fig. S4B). Allele-specific PCR analysis also confirmed the clonality of the three breast tumor cell lines (see Supplemental Fig. S4C–E). (C) RNA SNP6 analysis shows levels of X-linked gene allelic expression. X-linked genes known as subject to XCI (Carrel and Willard 2005; Cotton et al. 2013) and/or considered as monoallelically expressed in WI-38 clones (i.e., for each informative gene, <2/3 of the SNPs were observed as biallelically expressed) are shown on the boxplots. (D) RNA-seq analysis shows levels of X-linked gene allelic expression. X-linked shown on the boxplots are known to be subject to XCI (Carrel and Willard 2005; Cotton et al. 2013) and/or are considered as monoallelically expressed in WI-38 clones (i.e., for each informative gene, the allelic expression ratio is <40, i.e., expressed <20% on one of the two alleles). (E) Summary of the informative genes identified by the RNA SNP6 and RNA-seq approaches. Genes “known as subject to XCI” or “known to escape from XCI” refer to previous studies (Carrel and Willard 2005; Cotton et al. 2013). WI-38 data correspond to the two clones.











