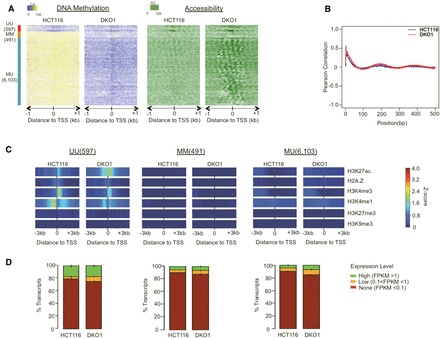
The loss of DNA methylation has little effect on the chromatin structure of non-CGI promoters. (A) NOMe-seq methylation levels were aligned to 7191 TSSs for annotated non-CGI promoters and displayed as in Figure 2. (B) Pearson autocorrelation of GC (accessibility) methylation levels was calculated independently for each cell type and displayed as in Figure 2. (C) Within each promoter class, the Z-score enrichment level of each histone mark is displayed as in Figure 2. (D) FPKM transcript values for all genes were divided into three levels and displayed as in Figure 2.











