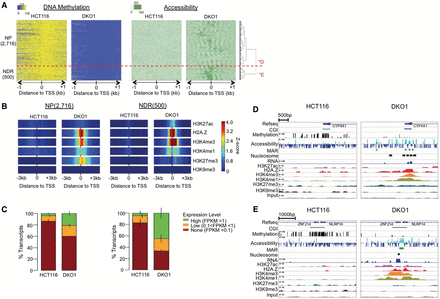
CGI promoters losing methylation acquire either an active or poised chromatin state, each with a distinct nucleosomal profile and histone modifications that resemble their respective patterns in normal colon cells. (A) NOMe-seq reads were aligned to 3216 MU CGI TSS from Figure 2 and hierarchically clustered based on the accessibility level in DKO1 cells. Two top-level clusters were detected in the unsupervised clustering: NP gained nucleosome positioning alone, while NDR promoters also gained nucleosome depletion. (B) Enrichment of each histone mark for the two clusters, displayed as in Figure 2. (C) Transcript level for each promoter class as displayed in Figure 2. (D) IGV browser view of NP gene CYP4X1 and (E) NDR gene ZNF214 (labeled as *D and *E in A). IGV views show DNA methylation (CG) and accessibility (GC) NOMe-seq methylation levels and ChIP-seq histone marks, with HCT116 and DKO1 plotted side by side. Methyltransferase accessible regions (MARs) and mononucleosomes were identified by our hidden Markov model (HMM), as described in Methods.











