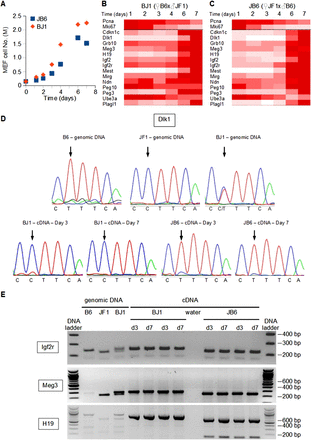
Monoallelic, parent-of-origin-dependent expression of imprinted genes is not altered in quiescent versus proliferating mouse embryonic fibroblasts. (A) Time course of cell numbers. JB6 and BJ1 MEFs were grown in vitro until they reached confluence. (B,C) Expression levels of proliferation markers (Pcna, Mki67) and representative IGs were monitored by real-time PCR in JB6 (B) and BJ1 (C) MEFs. Data for each gene are expressed as the percentage of the maximal expression levels for that gene. (D) Sequence of a polymorphic region (C[T/C]TTCA) of the Dlk1 gene (top panels) and transcripts (bottom panels). Genomic DNAs from C57BL/6J (B6), JF1/Ms, and C57BL/6J × JF1/Ms (BJ1) were sequenced and display the T, C, and C/T alleles, respectively. Sequencing of cDNAs from proliferating (day 3) and quiescent (day 7) MEFs derived from C57BL/6J × JF1/Ms (BJ1) and JF1/Ms × C57BL/6J (JB6) crosses indicates that only the paternal Dlk1 allele is expressed. (E) Genomic DNA and cDNAs were PCR amplified at the indicated loci from the indicated crosses. The amplicons from the Meg3 and H19 loci were digested with Bsh1236I and BclI, respectively. Digested and undigested amplicons were run on an ethidium bromide–stained agarose gel.











