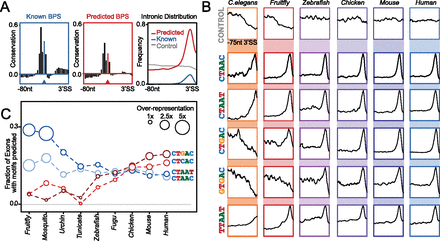
Branchpoint prediction and usage in different species. (A) Vertebrate conservation profile (100 species) of known (blue, left panel) and predicted (red, middle panel) branchpoint sequence (BPS) motifs. Frequency distribution (right panel) of known (blue) and predicted (red) motifs relative to the 3′ splice site, with matched scrambled control (gray) indicated. (B) Frequency distribution of common branchpoint motifs across a 75-nt window upstream of the 3′ splice site from the range of model organism genomes reveals differential usage of branch motifs across different lineages. (C) Bubble plot/histogram indicates the proportion of exons from model organism genomes that are associated with selected motifs. Circle radius is proportional to motif overrepresentation against the expected background in each organism.











