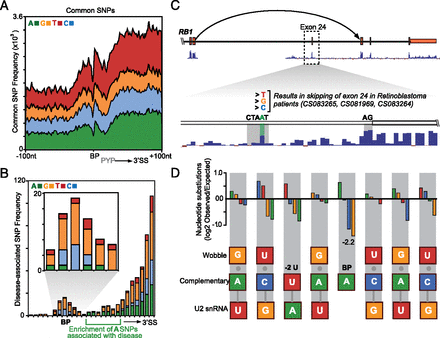
Common and disease-associated genetic variation at branchpoints. (A) Frequency distribution of common SNPs across a 200-nt window shows that branchpoints are refractory to common genetic variation (note that downstream exon enrichment is confounded by SNP ascertainment bias). (B) Frequency distribution of disease-associated SNPs (with inset showing detail) indicates enrichment at branchpoint. (C) UCSC Genome Browser view of nucleotide identified as a branchpoint (A; green) in the Retinoblastoma gene (RB1). Mutations at this nucleotide have been associated with exon 24 skipping (dashed arrow) in retinoblastoma patients (Houdayer et al. 2008; Zhang et al. 2008). High conservation (Vertebrate 46way) of branchpoint indicated by lower blue histogram. (D) Histogram indicates fold enrichment for nucleotide substitutions (SNPs) between individual human genomes. Fold enrichment indicates the observed rate of SNPs relative to the expected genome background rate of nucleotide substitution. This demonstrates enrichment for SNPs that maintain complementary or wobble binding between the B-box and U2 snRNA. U2 snRNA and B-box base-pairing nucleotide possibilities are indicated in the lower schematic.











