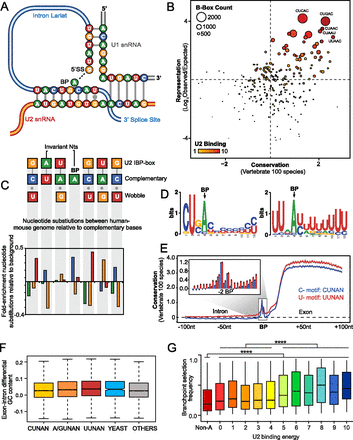
Branchpoint motifs and conservation. (A) Schematic illustrating base-pairing of consensus branchpoint flanking sequence to U2 snRNA IBP-box. (B) Bubble plot indicating the overrepresentation and conservation of pentamer sequences overlapping branchpoints. Color indicates predicted U2 binding strength and circle radius is proportional to total motif count. Overrepresented motifs, such as CUGAC, exhibit both high conservation and predicted U2 binding strength. (C) Enrichment for substitutions that maintain U2:B-box binding. The upper schematic shows the predominance of G and U bases within the U2 snRNA IBP box that can bind via complementary or wobble base-pairing to two different possible opposing nucleotides in the B-box. The lower histogram indicates the fold enrichment for nucleotide substitutions at syntenic bases within the B-box between the mouse and human genome. Fold enrichment is normalized for background rates of nucleotide substitutions between mouse and human genomes. We observe enrichment for nucleotide substitutions that maintain complementary or wobble base-pairing between the B-box and U2 snRNA. (D) Example motifs identified de novo in sequences flanking the branchpoint (branchpoint at +4 nt). (E) Average nucleotide conservation score (phyloP 100 vertebrates) for 100 nucleotides flanking the branchpoints. U motifs indicated in red; C-motif indicated in blue. Canonical CUAAC motif is not included in order to examine only derived motifs. Shaded areas represent the 95% confidence intervals. (Inset) Twenty nucleotides around the branchpoint, error bars are 95% confidence intervals. (F) Box-whisker plot (5%–95% range) of GC% differential between branchpoint introns and associated downstream exons for various families of B-box elements: (Yeast) CUAAC; (others) motifs without a branchpoint adenosine. (G) Box-whisker plot (2.5–97.5% range) of relationship between U2 binding energy and branchpoint selection frequency (per exon). (Non A) Nonadenosine branchpoints. Summary of significant differences shown. (****) P < 0.0001; one-way ANOVA with Tukey correction for multiple testing.











