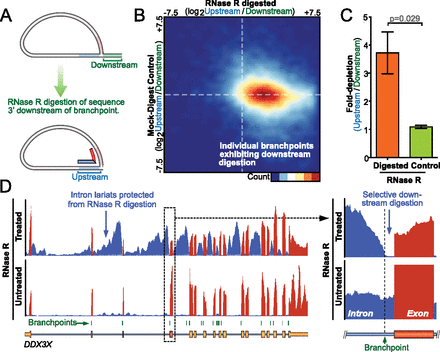
RNase R digestion of intron lariats. (A) Schematic showing digestion of the lariat tail (green) downstream from the branchpoint by RNase R, while the closed lariat protects sequences upstream of the branchpoint (blue). (B) Heatmap indicates the relative enrichment of sequences upstream of individual branchpoints within RNase R digested libraries (x-axis) compared to untreated control (y-axis) libraries. The majority of branchpoints show enrichment of upstream sequences after digestion (indicated by positive horizontal spread), whereas the clustering at zero on the y-axis indicates that sequences around individual branchpoints show no enrichment in mock-digested libraries. (C) Histogram indicates significant depletion of intronic sequences downstream from branchpoints (to the 3′SS) relative to the upstream region following RNase R digestion. Two human cell types, each with two biological replicates with matched untreated controls (paired t-test, n = 4, error bars SEM). (D) Genome Browser view showing enrichment of DDX3X intron lariats following RNase R digestion. Close detail indicates selective digestion of intronic sequence downstream from the identified branchpoint. In contrast, equivalent pre-mRNA is observed both up- and downstream from branchpoints in untreated libraries.











