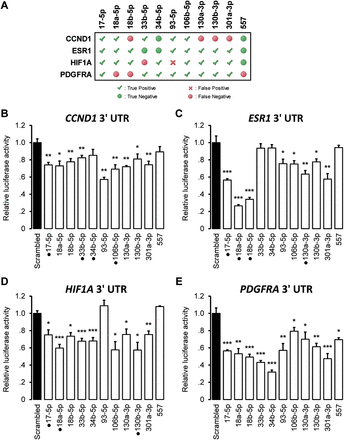
Regulatory potential of miRNA mediators. Multiple ceRNA interactions in a network including CCND1, ESR1, HIF1A, and PDGFRA (Fig. 4) were predicted to be mediated by common miRNAs. We tested the regulatory potential of 10 of these miRNAs biochemically, with miR-557 selected as a negative control. (A) Predicted miRNA–target interactions and a summary of biochemical validation, depicting true-positive, true-negative, false-positive, and false-negative predictions; down-regulation of 3′ UTR luciferase activity in response to miRNA-mimic transfection at P < 0.05 was taken as evidence for regulation. Luciferase activity after miRNA mimic transfection relative to transfection of scrambled control is shown for (B) CCND1, (C) ESR1, (D) HIF1A, and (E) PDGFRA 3′ UTRs. Punctuated mimics, for example, miR-17-5p for CCND1, ESR1, HIF1A, but not PDGFRA, correspond to previously validated interactions. Data are represented as mean ± SEM; (*) P < 0.05, (**) P < 0.01, (***) P < 0.001.











