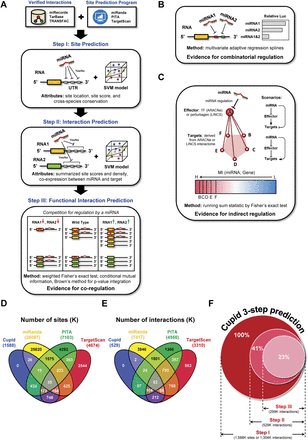
Methodology. (A) Cupid first reevaluates sites predicted by TargetScan, miRanda, and PITA, selecting and rescoring each candidate site (Step I). Sites are used to select and score miRNA-target interactions (Step II), which are then examined for evidence for mediating ceRNA interactions (Step III). In addition, to support interaction prediction we considered (B) evidence for combinatorial regulation between miRNAs and (C) evidence for indirect regulation by miRNAs through effectors. (D) The majority of site predictions by TargetScan, miRanda, and PITA are exclusive to a single algorithm; for example, <10% of sites predicted by miRanda are also predicted by another method. (E) Cupid predicted 529K miRNA-target interactions in Step II, excluding 60% of Step I candidate interactions. As a result, it makes considerably fewer predictions than TargetScan, miRanda, and PITA. (F) Cupid Step III predictions include less than a quarter of Step I candidate interactions.











