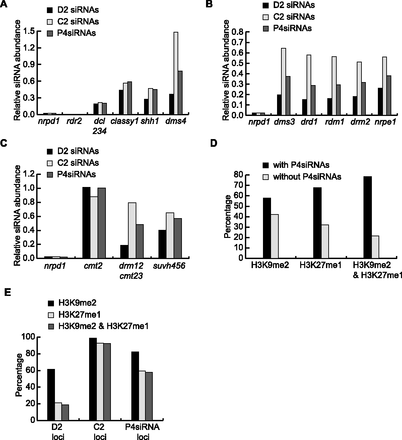
RdDM genes, epigenetic marks, and P4siRNA biogenesis. (A–C) Effects of mutations in various CHH methylation pathway genes on P4siRNA biogenesis. The relative abundance of D2, C2, and total P4siRNAs in various mutants compared to WT is shown. The analysis was performed with published sRNA-seq data (Lee et al. 2012; Law et al. 2013; Stroud et al. 2014). For each genotype, reads corresponding to P4siRNA loci were normalized against small RNAs from non-P4siRNA loci. P4siRNA loci were defined as those showing differentially expressed siRNAs between WT and nrpd1 (see Methods). A total of 47,742 total P4siRNA loci, 13,479 D2, and 19,039 C2 loci were used in the analysis. (A) Relative siRNA abundance in mutants in genes known to act in P4siRNA biogenesis. (B) Relative siRNA abundance in mutants in genes known to act downstream from P4siRNAs in RdDM. (C) Relative siRNA abundance in mutants in genes that confer CHH DNA methylation or histone H3K9 methylation. nrpd1 is included in B and C for comparison. (D,E) Overlap between P4siRNA loci and the epigenetic marks H3K9me2 or H3K27me1. Published ChIP-chip data were used to define regions with H3K9me2 or H3K27me1 (Roudier et al. 2011; Deleris et al. 2012). (D) Regions with H3K9me2, H3K27me1, or both H3K9me2 and H3K27me1 were divided into 500-bp windows. The number of windows where P4siRNAs were present or not was counted, and the percentage of total windows is shown. (E) The percentage of P4siRNA loci with H3K9me2, H3K27me1, or both. The number of D2, C2, and total P4siRNA loci with H3K9me2, H3K27me1, or both marks was determined, and the percentage of these total loci is shown.











