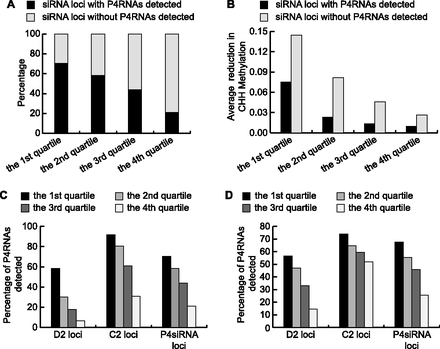
Decreased CHH DNA methylation in dcl234 compromises Pol IV transcription. (A–C) P4siRNA loci are divided into four quartiles according to P4siRNA abundance in WT, with the first quartile containing loci with the highest levels of P4siRNAs. (A) The percentage of P4siRNA loci with and without P4RNAs detected in our dsRNA-seq for the four quartiles. (B) The levels of CHH methylation decrease in dcl234 compared to WT for the four quartiles of P4siRNA loci with and without precursors detected. The decrease in CHH methylation was calculated using published methylome data (Stroud et al. 2013). (C) The percentage of P4RNAs detected at D2 and C2 siRNA loci for the four quartiles. Included in the analysis were 13,479 D2; 19,039 C2; and 47,742 total P4siRNA loci. (D) Correlation between P4RNA discovery and levels of CHH methylation at the siRNA loci. D2 and C2 siRNA loci are divided into four quartiles according to their CHH methylation levels in dcl234. The percentage of loci with P4RNAs detected in each quartile is shown. As CHH methylation levels decrease, the success rate of P4RNA discovery also decreases in total P4siRNA loci. In terms of P4RNA discovery, D2 loci are more sensitive to levels of CHH methylation.











