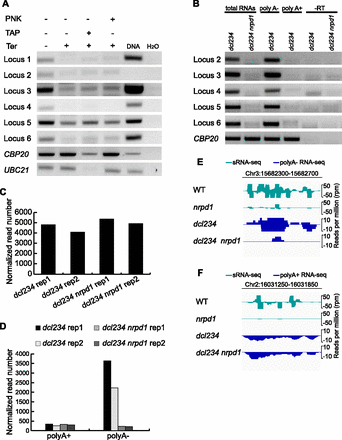
Features of P4RNAs. (A) Determination of the 5′ end structure of P4RNAs. Total RNAs from dcl234 were treated (+) or not (−) with various enzymes and subjected to random-primed RT-PCR to detect specific P4RNAs (loci 1–6) (Supplemental Table S1) with P4RNA-specific primers. (PNK) Polynucleotide kinase; (TAP) tobacco acid pyrophosphatase; (Ter) Terminator exonuclease. PCRs with genomic DNA and H2O (no RNAs in the reactions) were included as positive and negative controls, respectively. Transcripts from two genes, CBP20 and UBC21, were also detected by RT-PCR as controls. As expected, the levels of these RNAs were only reduced by digestion with both TAP and Ter. (B) Determination of the 3′ end structure of P4RNAs. Random-primed RT-PCR was performed on total RNAs from dcl234 and dcl234 nrpd1, and poly(A)-enriched and poly(A)-depleted RNAs from dcl234 to detect specific P4RNAs. (-RT) Reverse transcription was performed in the absence of reverse transcriptase. CBP20 served as a positive control for poly(A)+ RNAs. The CBP20 RT-PCR products in the poly(A)− fraction probably reflected degradation intermediates. (C) Abundance of reads at 1639 P4RNA regions with detectable transcripts in poly(A)+ RNA-seq. Two replicates of RNA-seq were conducted and the sum of the numbers of normalized reads is shown. (D) Abundance of transcripts at 698 P4RNA regions discovered through the initial RNA-seq, as determined by RNA-seq from poly(A)+ and poly(A)− RNAs. The reduction in transcript abundance in dcl234 nrpd1 was only observed in poly(A)− RNAs, indicating that P4RNAs lack poly(A) tails. (E) A genome browser view of reads at a P4siRNA locus on Chromosome 3 from sRNA-seq and poly(A)− RNA-seq. Read abundance is shown for both the Watson (top) and Crick (bottom) strands. (F) A genome browser view of reads at a P4siRNA locus on Chromosome 2 from sRNA-seq and poly(A)+ RNA-seq. Read abundance is shown for both the Watson (top) and Crick (bottom) strands.











