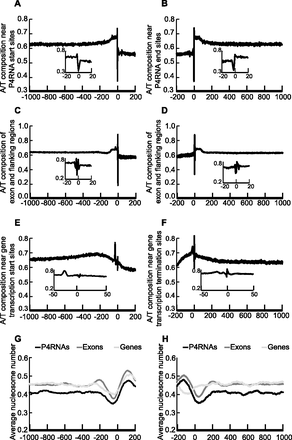
Genomic features of P4RNA and surrounding regions. A/T composition (A–F) and nucleosome occupancy (G–H) were examined at P4RNAs, exons, and genes. Exons and genes were according to TAIR10 annotation. Position 0 refers to the start site of transcription (for P4RNAs and genes) or the beginning of exons (in A, C, E, G) or the end site of transcription/end of exons (in B, D, F, H). Nucleotide positions upstream of and downstream from position 0 are represented by negative and positive numbers, respectively. Sequences were aligned at position 0 and the proportion of A/T nucleotides at each position is shown in A–F. (A,B) The A/T composition near the P4RNA start sites (A) or end sites (B). (C,D) The A/T composition of exons and flanking regions. (E,F) The A/T composition near protein-coding gene transcription start sites (E) or termination sites (F). In A–F, the insets display close-up views near position 0. (G,H) Average nucleosome occupancy near the start sites (G) or end sites (H) of P4RNAs, exons, and genes. The nucleosome positions were derived from published data (Chodavarapu et al. 2010).











