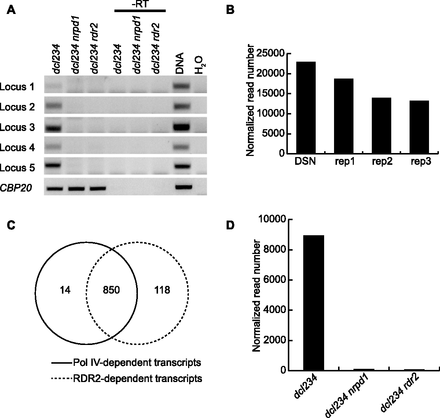
RDR2 has a similar effect as Pol IV on the abundance of P4RNAs. (A) Detection of P4RNAs by RT-PCR. Random-primed RT-PCR was performed on dcl234, dcl234 nrpd1, and dcl234 rdr2 to detect P4RNAs from five loci (Supplemental Table S1). PCRs with genomic DNA and H2O (no RNAs in the reactions) were included as positive and negative controls, respectively. (-RT) Reverse transcription was performed in the absence of reverse transcriptase. CBP20, a genic transcript, was included as a loading control. (B) DSN normalization moderately enriched the coverage of reads at P4RNA loci by RNA-seq. The total numbers of normalized reads at 47,442 P4siRNA loci from one replicate of dcl234 RNA-seq-DSN and three replicates of dcl234 RNA-seq are shown. (C) Venn diagram showing the overlap between regions with Pol IV-dependent transcripts and regions with RDR2-dependent transcripts as determined by RNA-seq-DSN of dcl234, dcl234 nrpd1, and dcl234 rdr2. (D) Abundance of Pol IV- and RDR2-dependent RNAs at the 850 Pol IV- and RDR2-dependent loci in C. The total numbers of normalized reads at these loci in RNA-seq-DSN are shown.











