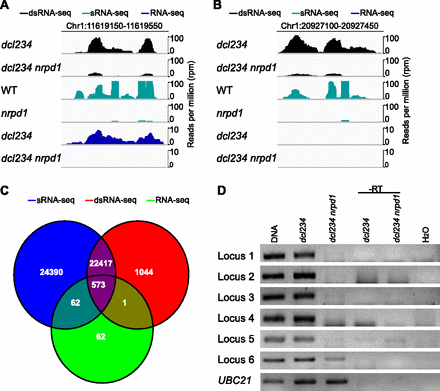
Genome-wide discovery of P4RNAs as P4siRNA precursors. (A,B) Genome browser views of small RNA reads and P4RNA reads at two representative P4siRNA loci. The read counts (in reads per million [RPM]) include reads from both strands. The top two, middle two, and bottom two rows represent reads from dsRNA-seq, sRNA-seq, and RNA-seq, respectively. In A, P4RNAs were detected by both dsRNA-seq and RNA-seq. In B, P4RNAs were only detected by dsRNA-seq. (C) Venn diagram showing the overlap of P4RNA regions discovered through dsRNA-seq or RNA-seq with P4siRNA regions discovered through sRNA-seq. Note that dsRNA-seq and RNA-seq were conducted with dcl234 and dcl234 nrpd1, and sRNA-seq was conducted with WT and nrpd1. (D) Random-primed RT-PCR analysis of P4RNAs discovered through RNA-seq on RNA samples from dcl234 and dcl234 nrpd1. Genomic DNA was included as the positive control for the PCR. Reverse transcriptase (-RT) was omitted from the reverse transcription reactions. “-RT” and H2O (no RNAs in the reactions) served as negative controls. The genomic locations of the loci can be found in Supplemental Table S1.











