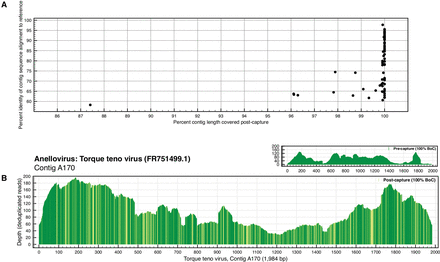
Targeted sequence capture identifies divergent sequences. (A) The percentage identity of the top high-scoring segment pair (HSP) identified from the BLAST alignment of anellovirus contig sequences to the references used to design ViroCap is plotted on the y-axis. The x-axis represents the percentage of the length of the anellovirus contig covered after targeted sequence capture. (B) This coverage plot represents the sequence coverage of a divergent anellovirus contig sequence. The figure is designed as described in the figure legend for Figure 2, with the following addition: The post-capture coverage plot is shaded to show regions of nucleotide sequence variation between the anellovirus contig and the most similar reference genome in the ViroCap panel. Dark shading represents areas of identical sequence, and each position with nucleotide mismatch between aligned sequences is shown in the lighter color. All of the HSPs are shown, rather than just the top HSP.











