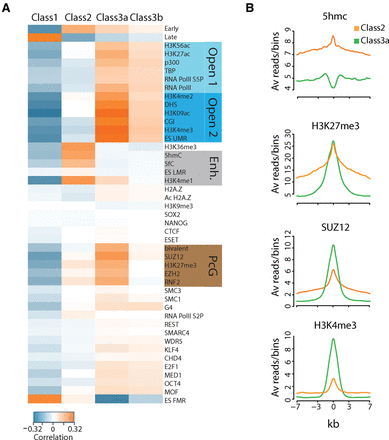
Origin classes are linked to specific chromatin signatures. (A) Association of chromatin marks/factors within each class. Pearson correlation between pairs of chromatin marks/factors for all IS in each class. Class 1 IS (n = 28,443) are correlated with closed chromatin marks and anticorrelated with all other marks/factors. Class 2 IS (n = 14,873) are the only IS positively correlated with enhancer features (Enh.), whereas Class 3a and 3b IS (n = 10,547) are associated with Open 2 and Polycomb group (PcG) chromatin marks/factors. (B) ChIP-seq signals for 5hmC, H3K27me3, SUZ12, and H3K4me3 around ±7 kb from the center of Class 2 and Class 3a IS.











