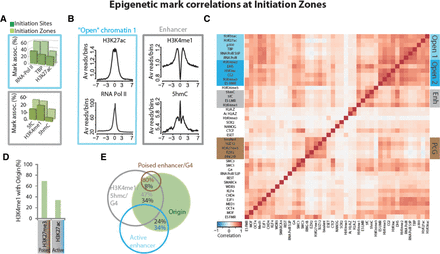
Epigenetic marks and chromatin factors at IZ. (A,B) Specific chromatin marks/factors associated with open chromatin-1 (light blue boxes) and enhancers (gray boxes) at initiation zones. (A) Distribution of chromatin marks/factors relative to IS (dark green) and IZ (light green). (B) ChIP-seq signals for H3K27ac, RNA Pol II, H3K4me1, and 5hmC around ±7 kb from the IS. (C) Hierarchical clustering of Pearson correlations between pairs of marks and/or chromatin factors at IZ. Marks and factors in the heatmap are ordered according to the clustering performed for the IS (see Fig. 2C). Positive correlations between pairs of marks/factors in individual IZ regions are symbolized by a gradation of red. Negative correlations are symbolized by a gradation of blue. (D) Percent overlap between poised (H3K4me1/H3K27me3) and active (H3K4me1/H3K27ac) enhancers within IZ. (E) Overlap between marks/factors and origins.











