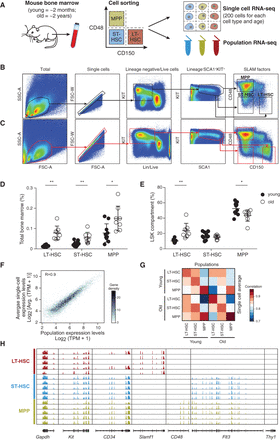
Single-cell RNA-seq of young and old HSCs. (A) Overview of experimental design. (B,C) Sorting strategy for isolating LT-HSCs (LSK CD48−CD150+), ST-HSCs (LSK CD48−CD150−), and MPPs (LSK CD48+CD150−) from young (B) and old (C) C57BL/6 mice. (D,E) LT-HSC compartment expands during aging. Shown are frequencies of LT-HSC, ST-HSC, and MPPs (x-axis) in young (black) and old (white) C57BL/6 mice as a percentage of bone marrow (BM; D) or stem cell compartment (lineage− SCA1+KIT+, LSK; E). Statistically significant differences are as follows: (**) P < 0.01, (*) P < 0.05; n = 8–10. (F) Single-cell RNA-seq recapitulates population RNA-seq. Shown are expression levels for all genes calculated from RNA-seq of a population of young LT-HSCs (x-axis) and by averaging expression levels from approximately 200 single young LT-HSCs (y-axis). The Pearson correlation coefficient (r = 0.9) is denoted. Gray scale bar indicates gene density. (G) Heatmap of Pearson correlation coefficients (r; color bar) between pairs of RNA-seq profiles of populations (columns) and matching averaged single-cell data (rows) from C57BL/6. (H) RNA-seq coverage of known cell surface markers in representative cells from young C57BL/6 mice (plot generated by the Integrative Genome Viewer 2.3) (Robinson et al. 2011; Thorvaldsdottir et al. 2013).











