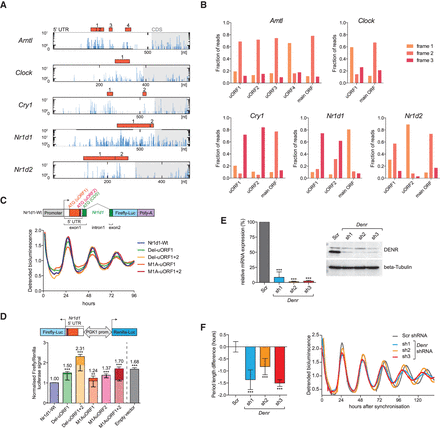
uORF translation is prevalent in core clock transcripts and impacts clock functions. (A) The 5′ UTRs of the depicted clock transcripts all contained at least one translated AUG-initiated uORF. Distribution of raw read counts of RPF reads (blue) along the 5′ UTR (white region in box) and the first 200 nt of the CDS (gray shaded region) is shown for the timepoint with maximal CDS translation. Red filled boxes indicate AUG-initiated uORFs within the 5′ UTR. Predicted uORFs for each gene are serially numbered. (B) Frame preference of uORF-mapping footprints. The fractions of footprints aligning to the three reading frames are shown for the uORFs shown in A and for the main ORF (CDS). Frame definition is relative to the annotated 5′ end of the transcript; please note that frame definition is different from that in Figure 1F. Most uORFs are thus covered by footprints that have a similar degree of frame preference as the main ORF-mapping footprints, indicating that uORF-mapping reads likely originate from processive translation. (C, top) Schematic showing the wild-type Nr1d1-firefly luciferase (FL) reporter gene consisting of a genomic Nr1d1 fragment in which FL (blue) is expressed in fusion with the first 15 amino acids of NR1D1 (green). In exon 1, the location of uORF1 and -2 (red) and their predicted start codons within the 5′ UTR is shown. (Bottom) Real-time bioluminescence recordings of luciferase rhythms in NIH3T3 cells lentivirally transduced with the Nr1d1-FL reporter (Wt) and various mutants in which either uORFs are deleted (Del mutants) or uORF initiation codons are mutated to an alanine codon (M1A mutants). Cells were synchronized with dexamethasone. Raw bioluminescence was detrended using a 24-h moving average, and one representative replicate of a total N = 3–9 is shown. (D, top) Schematic representation of the dual luciferase reporter construct to measure how the Nr1d1 5′ UTR (Wt/mutants) influences the expression of the FL CDS. From the same bidirectional promoter, Renilla luciferase (RL) is expressed for internal control. (Bottom) Results of dual luciferase assay where FL signals were internally normalized to RL. Empty vector (gray) only contained the vector-encoded 5′ UTR. Experiments were performed in NIH3T3 cells (N = 2–4 independent experiments of triplicates). Lighter shading of the bars indicates the proportion of the increase that can be attributed to increased FL mRNA abundance in the mutants (measured by qRT-PCR), leaving the remainder of the increase (darker shading) attributable to translation. Note that whenever the translated uORF1 and -2 of the Nr1d1 5′ UTR are deleted (Del mutants) or just the initiation codons are mutated to alanine codons (M1A mutants), the inhibitory activity of the Nr1d1 5′ UTR is relieved. uORF1 and uORF2 appear to have an additive inhibitory effect on main ORF translation (cf. M1A uORF1 and -2 single mutants with the double mutant). For a schematic of the mutants, see also Supplemental Figure S9C. (E, left) Relative mRNA levels (RT-qPCR; normalized to expression of control gene Nudt4) of Denr in DBP-Luciferase reporter–expressing NIH3T3 cells transduced with scramble shRNA (Scr; serving as control) or three different shRNAs targeting Denr (N = 3). All shRNAs reduced Denr expression to <10%. (Right) Western blot analysis for DENR indicated efficient depletion at the protein level. Beta tubulin served as loading control. (F) shRNA-mediated Denr knockdown causes a short period phenotype of free-running circadian rhythms in NIH3T3 cells. (Left) Summary of period change engendered by Denr shRNAs 1–3 relative to Scr (control) shRNA in cells expressing the DBP-Luciferase reporter (N = 3–8). (Right) Representative bioluminescence tracks of Scr (control) and Denr shRNA-transduced DBP-Luciferase cells. Depending on the shRNA, the period of free-running circadian oscillations was 0.8–1.5 h shorter. (D–F) Bar graphs, mean ± SD; (**) P < 0.01, (***) P < 0.001 (t-test). P-value in D refers to differences in FL/RL activities (darker shading).











